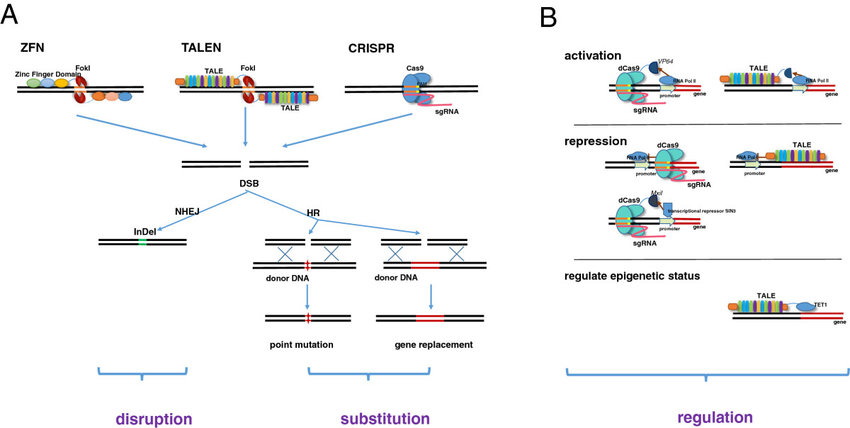
There are approximately 6-10,000 monogenic diseases in humans – diseases caused by a mutation in a single gene. For 95% of them there is no treatment, and there are few cures. This is partly why there was so much excitement when CRISPR technology was introduced in 2013 (the research goes back to 1987, but its use for genetic engineering was proposed in 2013, and first used in 2015). However, a competing gene editing technology, TALEN, was introduced in 2012, but has received much less public attention. Both technologies are powerful, but have different strengths and weaknesses, and likely will complement each other going forward.
CRISPR
I’m still trying to memorize what the acronym CRISPR stands for – Clustered Regularly Interspaced Short Palindromic Repeats (whew, did that without looking). This genetic sequence was discovered in bacteria, that use it as part of their immune defense. CRISPR has the ability to target specific sequences of DNA and to carry an enzyme as a payload. The most common enzyme paired with CRISPR today is Cas9, but there are others. Cas9 slices the DNA, which in its simplest application can silence the gene. However, this can also be paired with DNA repair mechanisms used to insert new strands of DNA and thereby perform gene editing.
The CRISPR Cas9 technology is evolving rapidly so it is difficult to say what the full potential of this technology is. It has already, however, been used to treat patients with sickle cell anemia and is being studied for cancer therapy and other applications. Obviously the potential is huge, even beyond medicine to agriculture, manufacturing, and environmental applications.
The big advantage of CRISPR is that it is cheap and easy (has high efficiency) – 3-6 fold cheaper than its competitor. This has significantly lowered the bar for laboratories and companies to get involved in gene-editing technology. This is already dramatically increasing the pace of gene-editing research, and will likely result in an explosion of genetic therapies, especially considering the unmet need of genetic disease. CRISPR can also be more easily multiplexed, with multiple targets tackled simultaneously.
The main limitation of CRISPR, however, is that it has limited specificity. Given a random stretch of DNA, CRISPR can find its target only about 70% of the time (70% efficiency). However, this is much greater than older methods. Further, there are many off-target effects where Cas9 will slice the DNA that is not targeted by the CRISPR. It is not clear how clinically significant these off-target cuts are, but it is certainly not desirable.
In addition to off-target effects, we can think of limitations in terms of mosaicism and allele heterogeneity. Mosaicism refers to the fact that not all the target cells will be affected. For example, if a specific population of cells in a mouse are targeted with CRISPR-Cas9, only some of them will have the edited genes. Depending on the disease being targeted, this can have a significant effect on the outcome. Further, when the cleaved DNA is repaired, it can be repaired in numerous ways, introducing random mutations or leading to incomplete introduction of the desired new DNA.
Again, CRISPR research is advancing quickly, and already researchers have identified ways to make the CRISPR reaction slower but more specific. It remains to be seen what an optimized CRISPR system will be.
TALEN
TALEN stands for Transcription activator-like effector nuclease and functions mainly through artificial restriction enzymes. These are enzymes that cleave DNA (like Cas9) but do so at a specific DNA sequence. Like the CRISPR-Cas9 system, TALENs have two domains, one that targets the DNA and one that cleaves it. The technology involves creating specific restriction enzymes that can target any desired sequence of DNA.
TALEN is more expensive and time-consuming than CRISPR – but to be clear, still much better than older methods. It is also less efficient that CRISPR, affecting about 1 in 140 target sequences. However, it has a big advantage – it is much more specific, with few off-target effects.
However, a recent study shows that under certain circumstances TALEN can be more efficient that CRISPR. In very compact parts of the genome, so-called heterochromatin, CRISPR efficiency goes way down. In a direct comparison the researchers found that TALEN was 5-6 times more efficient than CRISPR in effecting changes to heterochromatin DNA.
These differences suggest that both TALEN and CRISPR are likely to be used in the future as complementary technologies. For completeness, there is also a third option called Zinc Finger proteins (ZNFs) which is still considered one of the modern genetic engineering techniques, but it is older than TALEN and not as good, as far as I can tell, on any standard measure (efficiency, specificity, cost and time). It still may find a role going forward if it proves superior in certain applications. All three technologies are also still evolving.
While CRISPR, with good reason, has been getting all the popular press recently, there are several gene-editing technologies with different strengths and weaknesses that are likely to play a significant role in medicine in the future. Our ability to edit the genome quickly, cheaply, efficiently, and precisely is much greater than even just a decade ago. The last decade has been a clear inflection point, and going forward we can expect a dramatic increase in our ability to treat genetic diseases.
Even non-genetic diseases, like cancer, are potential targets for this technology. The early research using CRISPR as a form of anti-cancer targeted therapy is extremely encouraging.
At the same time, the proliferation of genetic engineering technology has to be carefully watched, with an eye toward regulation to prevent dangers and abuses. Like any advanced technology, genetic engineering can be both used for benefit or abused with unacceptable risks. Already a rogue researcher has used CRISPR to edit human embryos in an unethical way. But this is an entirely manageable issue as long as we both appreciate and respect the power of this technology.


